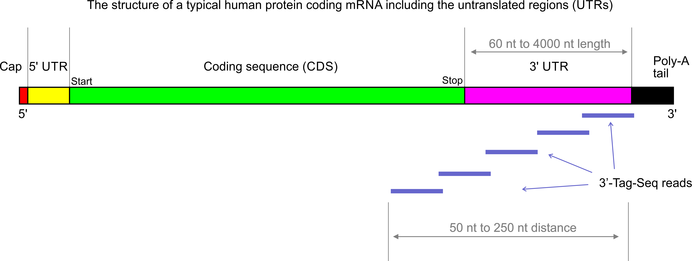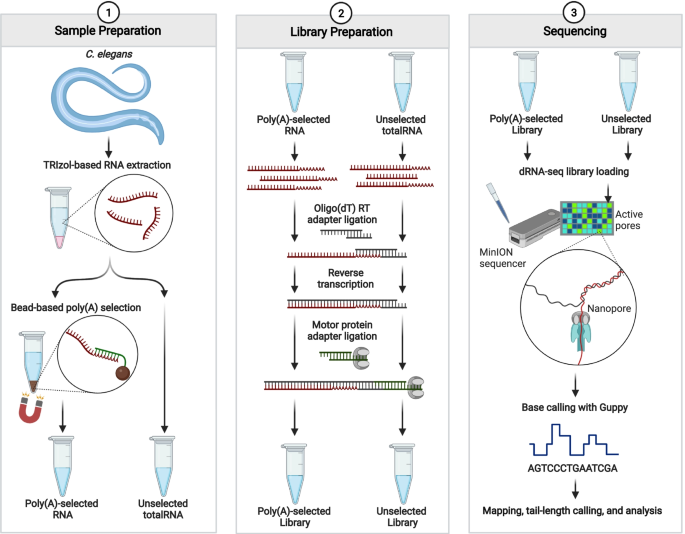
Poly(a) selection introduces bias and undue noise in direct RNA-sequencing | BMC Genomics | Full Text
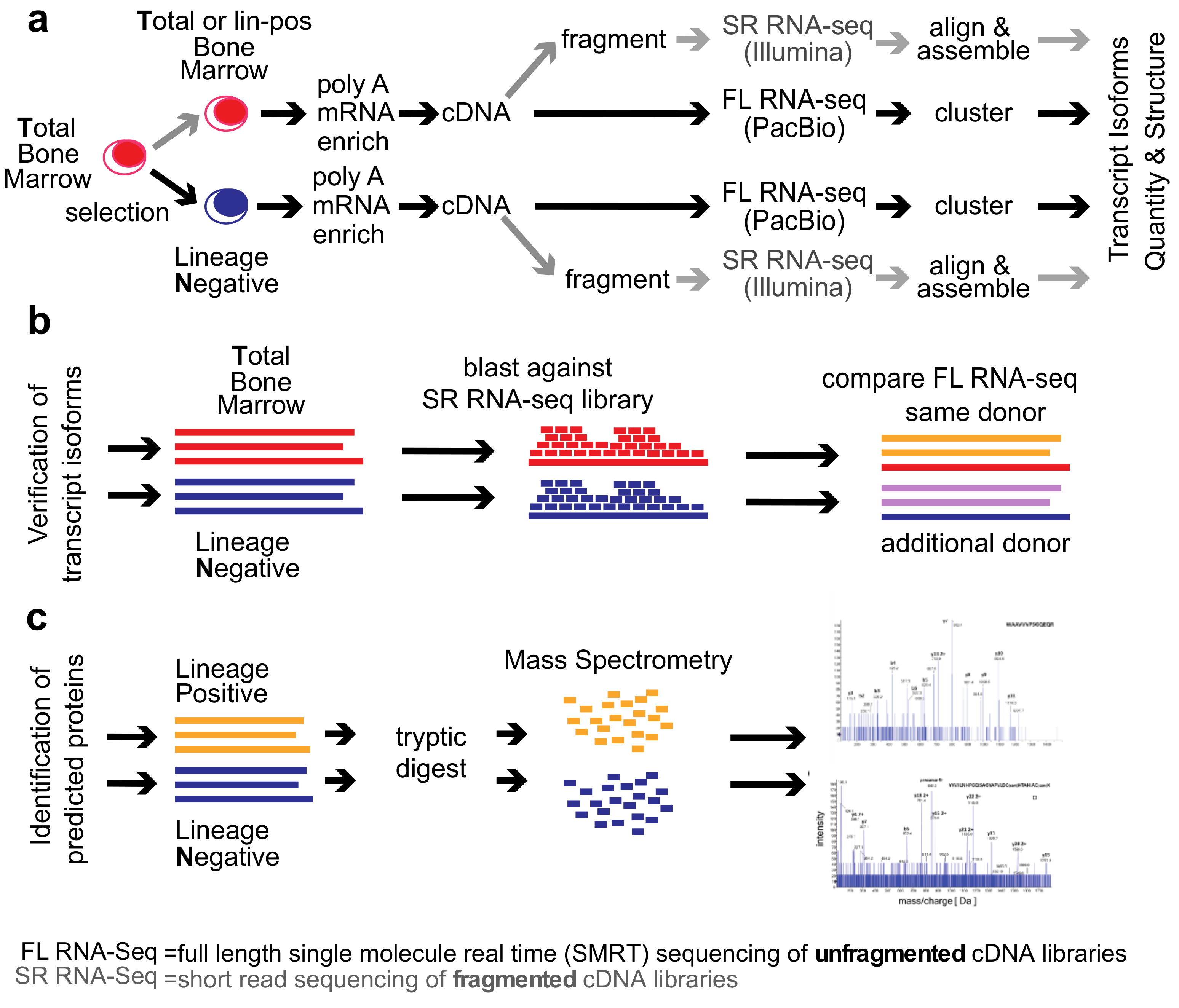
Genes | Free Full-Text | Single-Molecule Real-Time (SMRT) Full-Length RNA- Sequencing Reveals Novel and Distinct mRNA Isoforms in Human Bone Marrow Cell Subpopulations

Comparison of RNA-Seq by poly (A) capture, ribosomal RNA depletion, and DNA microarray for expression profiling | BMC Genomics | Full Text

Poly(A)-seq – A method for direct sequencing and analysis of the transcriptomic poly(A)-tails | RNA-Seq Blog

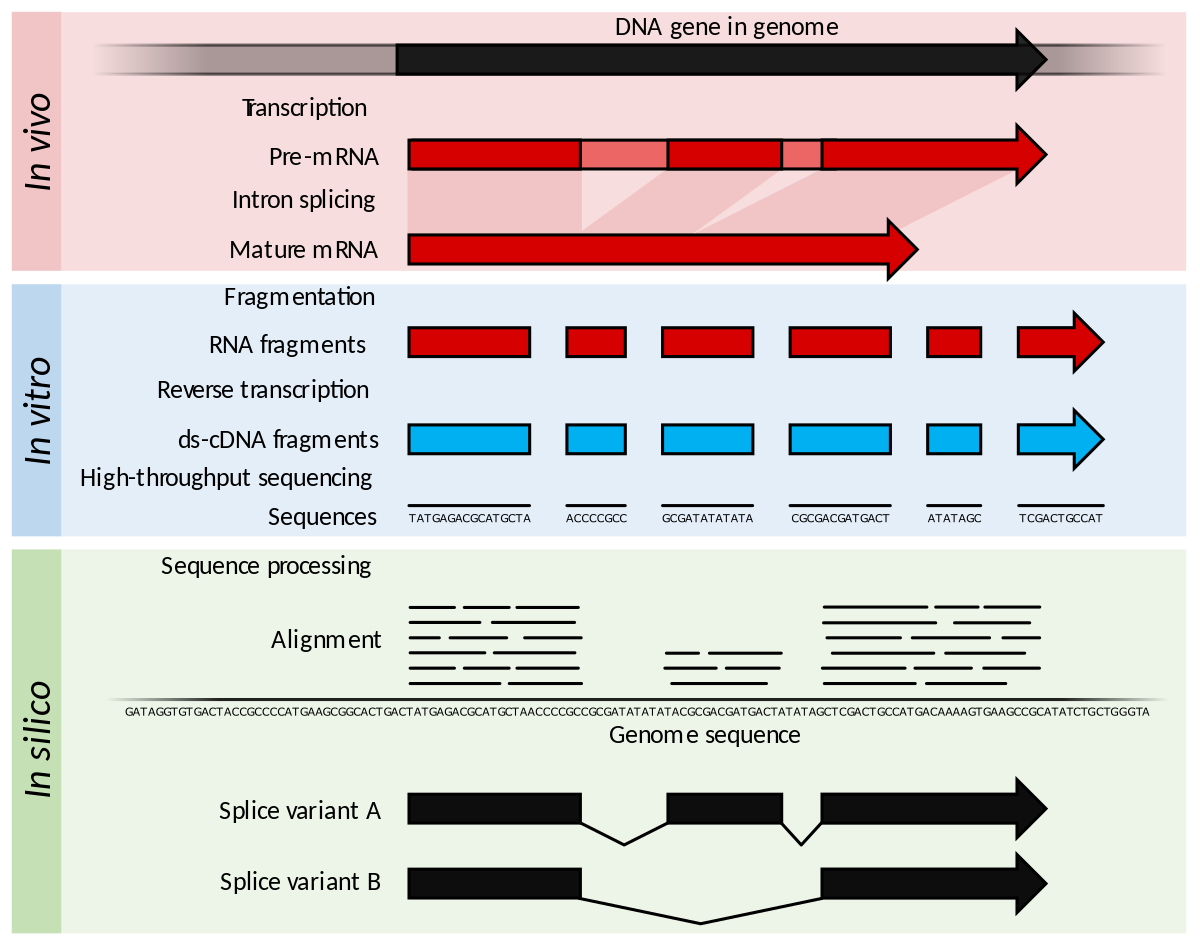
The workflow of the RNA-seq profiling of the nephron transcriptome.... | Download Scientific Diagram

Poly(A) capture full length cDNA sequencing improves the accuracy and detection ability of transcript quantification and alternative splicing events | Scientific Reports
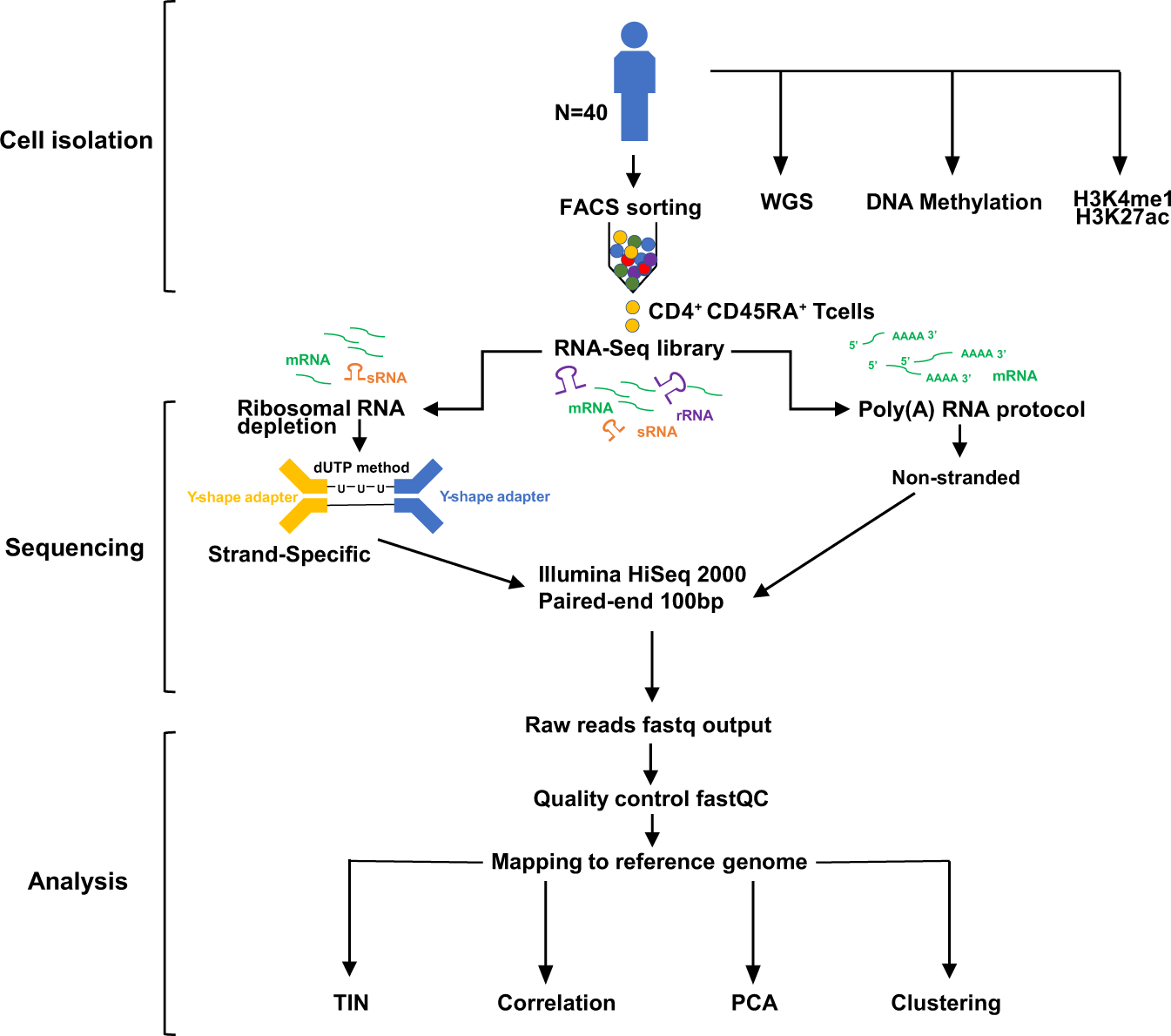
Paired rRNA-depleted and polyA-selected RNA sequencing data and supporting multi-omics data from human T cells | Scientific Data

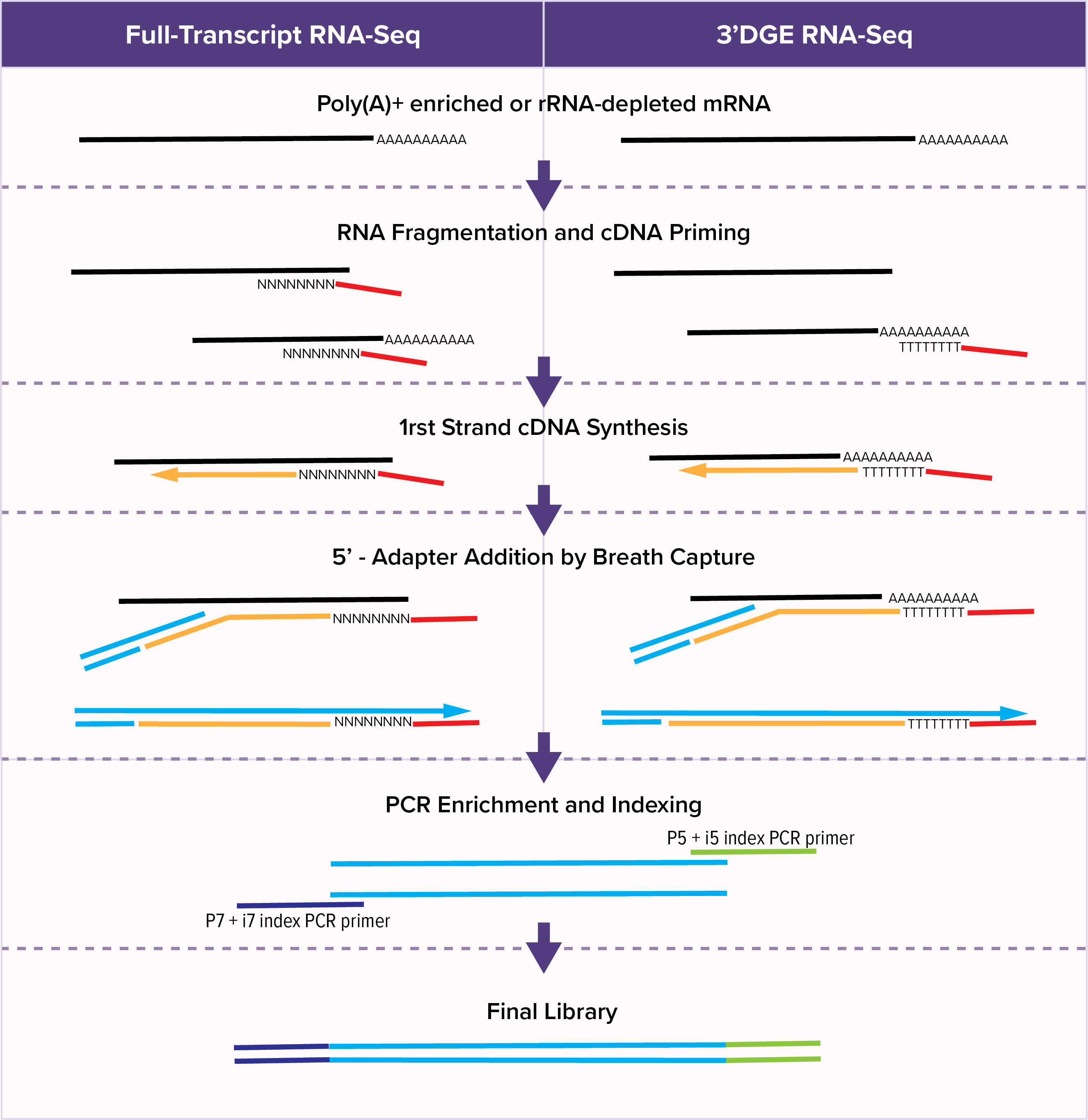
Diagram of LM-Seq sample preparation protocol. Poly-A-tailed mRNA is... | Download Scientific Diagram
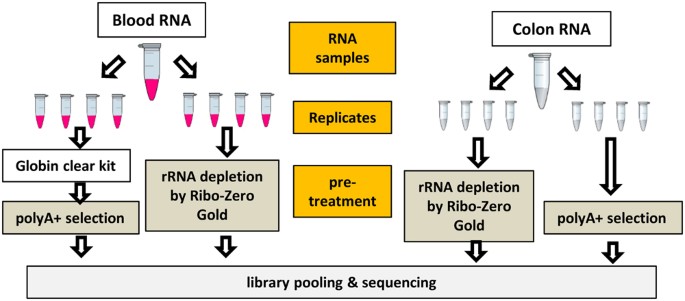
Evaluation of two main RNA-seq approaches for gene quantification in clinical RNA sequencing: polyA+ selection versus rRNA depletion | Scientific Reports
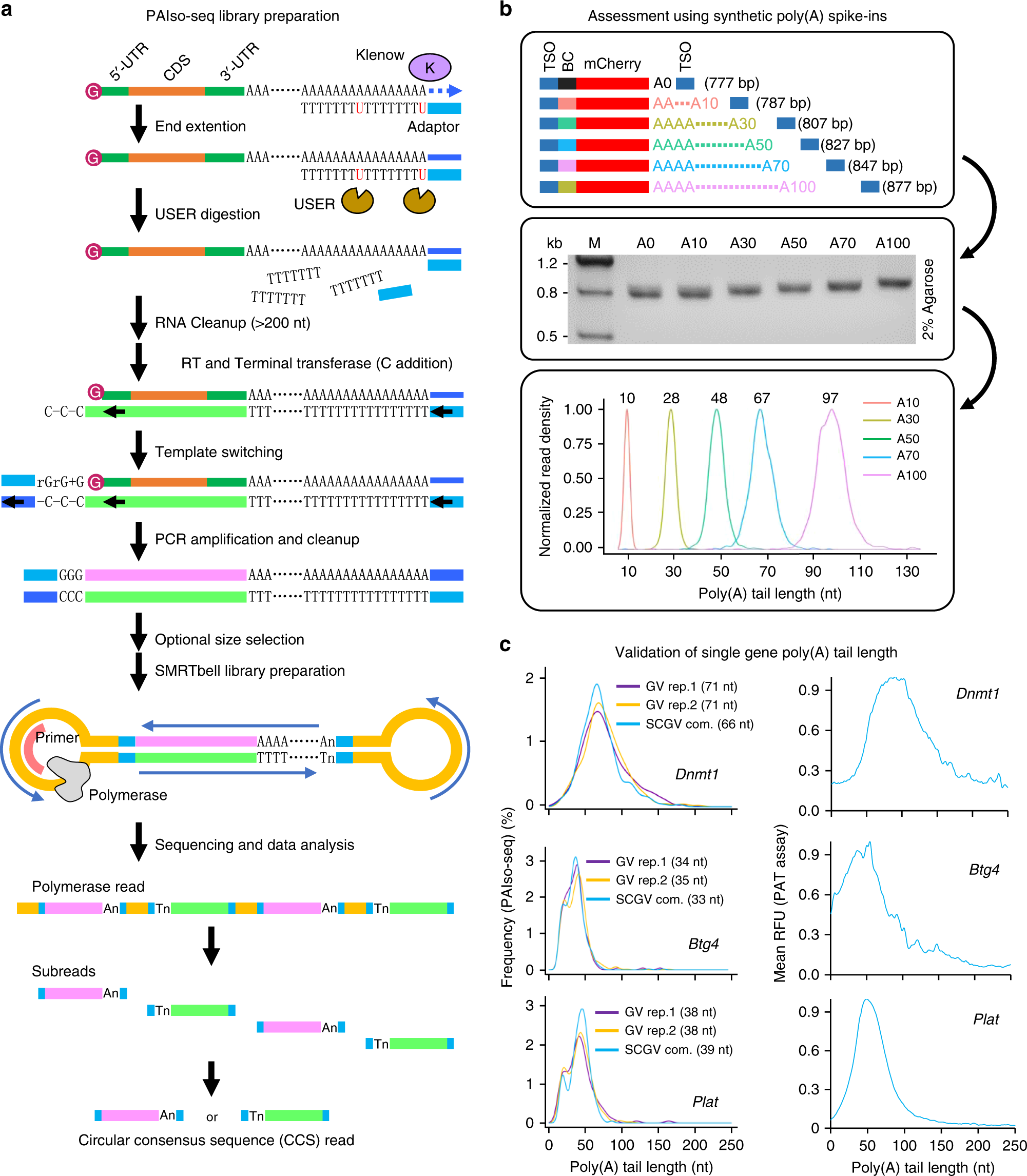
Poly(A) inclusive RNA isoform sequencing (PAIso−seq) reveals wide-spread non-adenosine residues within RNA poly(A) tails | Nature Communications
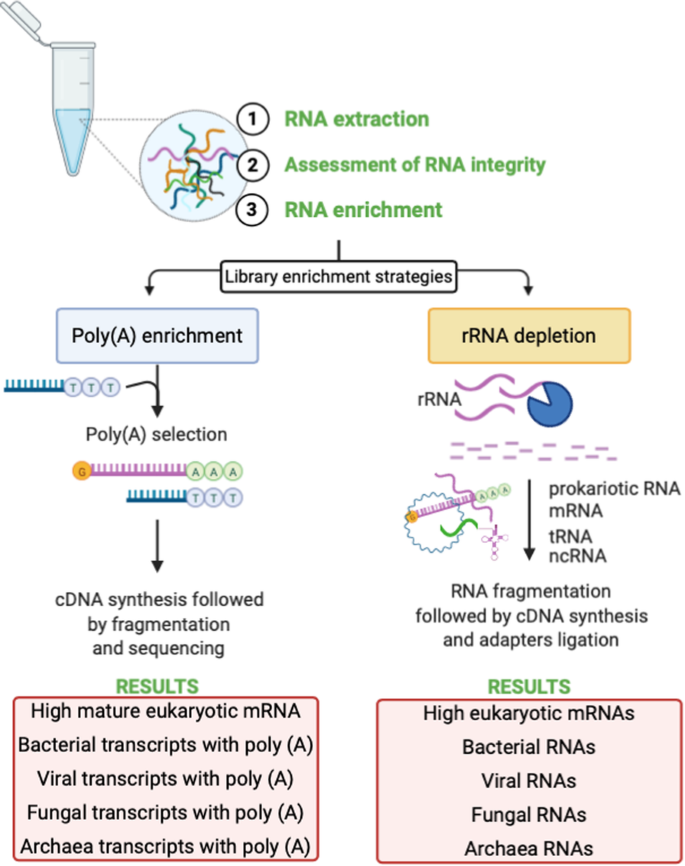
Omission of non-poly(A) viral transcripts from the tissue level atlas of the healthy human virome | BMC Biology | Full Text
Poly(A)-seq: A method for direct sequencing and analysis of the transcriptomic poly(A)-tails | PLOS ONE
Poly(A)-seq: A method for direct sequencing and analysis of the transcriptomic poly(A)-tails | PLOS ONE

Poly(A)-ClickSeq – click-chemistry for next-generation 3΄-end sequencing without RNA enrichment or fragmentation | RNA-Seq Blog

Multiple freeze-thaw cycles lead to a loss of consistency in poly(A)-enriched RNA sequencing | bioRxiv